The first-ever images of the protein complex that unwinds, splits, and copies double-stranded DNA reveal something rather different from the standard textbook view. The electron microscope images, created by scientists at the U.S. Department of Energy's Brookhaven National Laboratory with partners from Stony Brook University and Rockefeller University, offer new insight into how this molecular machinery functions, including new possibilities about its role in DNA "quality control" and cell differentiation. The images and implications are described in a paper published online by the journal Nature Structural & Molecular Biology, November 2, 2015.
"This work is a continuation of our long-standing research using electron microscopy to understand the mechanism of DNA replication, an essential function for every living cell," said Huilin Li, a biologist with a joint appointment at Brookhaven Lab and Stony Brook University.
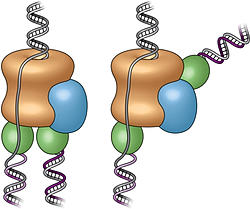
"These new images show the fully assembled and fully activated 'helicase' protein complex—which encircles and separates the two strands of the DNA double helix as it passes through a central pore in the structure—and how the helicase coordinates with the two 'polymerase' enzymes that duplicate each strand to copy the genome."
Three-dimensional structure of the active DNA helicase bound to the front-end DNA polymerase (Pol epsilon). The DNA polymerase epsilon (green) sits on top rather than the bottom of the helicase.
Studying this molecular machinery, known collectively as a "replisome," and the details of its DNA-copying process can help scientists understand what happens when DNA is miscopied—a major source of mutation that can lead to cancer—or learn more about how a single cell can eventually develop into the many cell types that make up a multicellular organism. But no one has produced a real structure of a replisome at any resolution for any organism until now.
"All the textbook drawings and descriptions of how a replisome should look and work are based on biochemical and genetic studies," Li said, likening the situation to the famous parable of the three blind men trying to describe an elephant, each looking at only one part. Those textbook drawings show the helicase moving along the DNA, separating the two strands of the double helix, with two polymerases located at the back where the DNA strand is split. In this configuration, the polymerases would add nucleotides (molecules containing the complementary A, T, G, and C bases of the genetic code) to the side-by-side split ends as they move out of the helicase to form two new complete double helix DNA strands.
Collaborating scientists and study coauthors Zuanning Yuan, a graduate student at Stony Brook University (standing), Huilin Li of Stony Brook and Brookhaven Lab (seated, back), and Jingchuan Sun of Brookhaven Lab (seated, front) examining protein structures.
To test these assumptions, Li's group turned to the technique they had previously used to study individual components of the helicase, electron microscopy (EM). Jingchuan Sun, an EM expert in Li's lab, was essential to the success of the work. He studied samples of replisomes from baker's yeast cells—a model for all nucleus-containing cells—prepared and provided by Roxana Georgescu in Michael O'Donnell's research group at Rockefeller University. O'Donnell's group had previously published biochemical results related to this work.
"DNA replication is one of the most fundamental processes of life, so it is every biochemist's dream to see what a replisome looks like," Sun said. "Our lab has expertise and a decade of experience using electron microscopy to study DNA replication, which has prepared us well to tackle the highly mobile therefore very challenging replisome structure. Working together with the O'Donnell lab, which has done beautiful functional studies on the yeast replisome, our two groups brought perfectly complementary expertise to this project," he said.

The team's first-ever images of an intact replisome revealed that only one of the polymerases is located at the back of the helicase. The other is on the front side of the helicase, where the helicase first encounters the double-stranded helix. This means that while one of the two split DNA strands is acted on by the polymerase at the back end, the other has to thread itself back through or around the helicase to reach the front-side polymerase before having its new complementary strand assembled.
The scientists were so surprised by this finding that they asked another group at Rockefeller, led by Brian Chait, to perform additional structural studies using mass spectrometry. Yi Shi, a postdoctoral fellow in Chait's group performed this work, which confirmed the electron-microscopy-based conclusion about the unexpected architecture of the replisome.
The counterintuitive position of one polymerase at the front of the helicase suggests that it may have an unforeseen function. The authors suggest several possibilities including keeping the two "daughter" strands separate to help organize them during replication and cell division. It might also be possible that, as the single strand moves over other portions of the structure, some "surveillance" protein components check for lesions or mistakes in the nucleotide sequence before it gets copied—a sort of molecular quality control.
This architecture could also potentially play an important role in developmental biology by providing a pathway for treating the two daughter strands differently. Many modifications to DNA, including how it is packaged with other proteins, control which of the many genes in the sequence are eventually expressed in cells. An asymmetric replisome may result in asymmetric treatment of the two daughter strands during cell division—an essential step for making different tissues within a multicellular organism.
As the paper concludes, "Clearly, further studies will be required to understand the functional implications of the unexpected replisome architecture reported here."
This research was funded by the National Institutes of Health (GM103314, GM109824, GM74985, AG29979, and GM38839) and by the Howard Hughes Medical Institute.


No comments:
Post a Comment